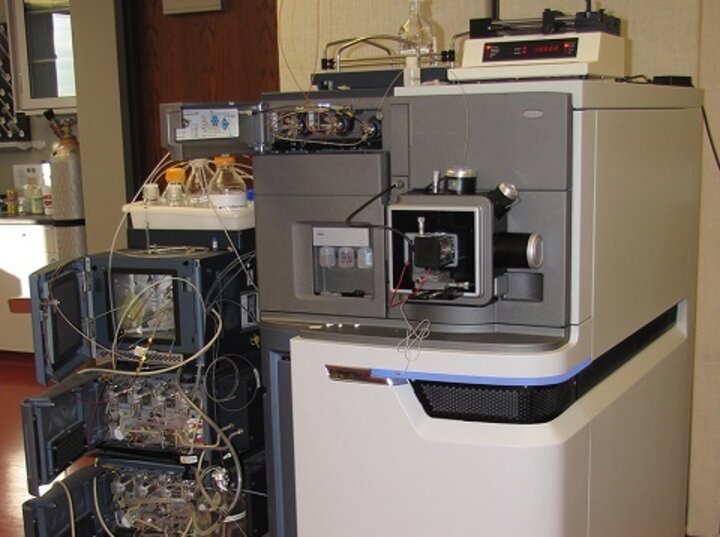
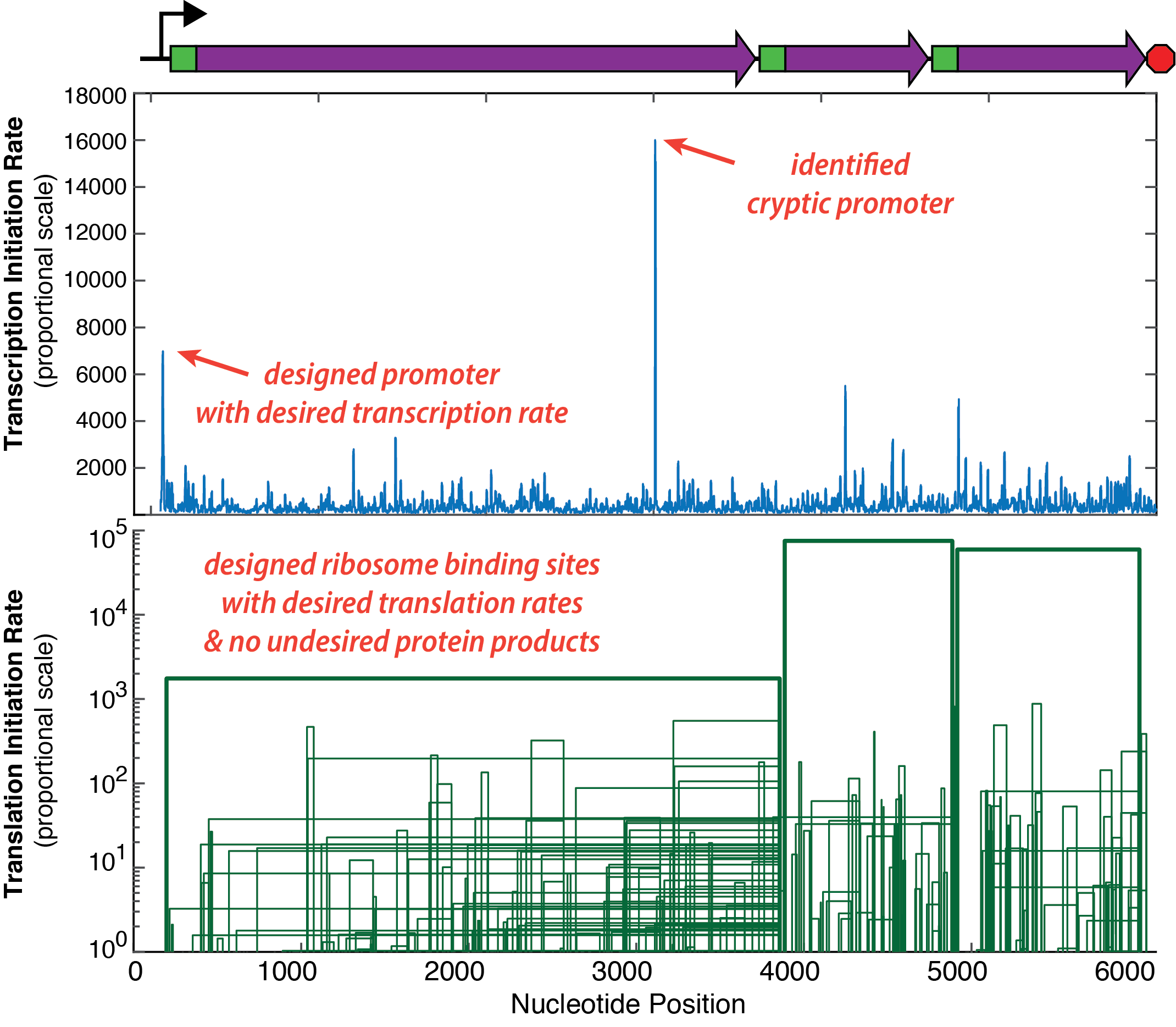
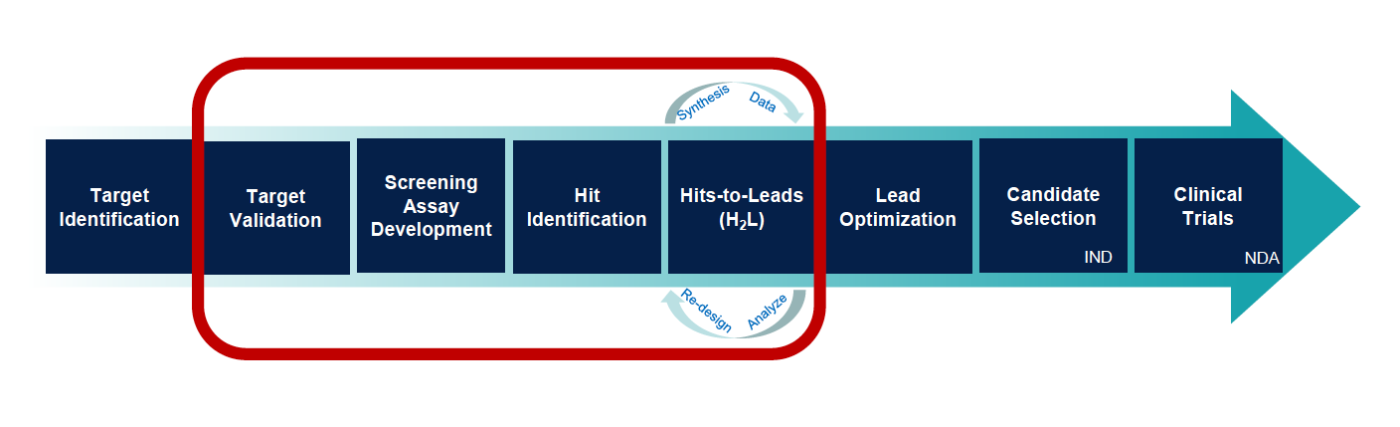
Introduction A Programmable Cloud Laboratory (PCL) is a laboratory that carries out experiments for researchers working at other institutions, using a web interface or API to enable automation and programmability. Past PCL efforts focused on creating highly automated laboratories that offered a small number of streamlined workflows, which limited their impact, as they are in direct competition with the thousands of existing biotech service providers – biofoundries, university core facilities, commercial services – that routinely carry out many thousands of high-quality workflows. However, existing service providers do not have the ability to interconnect their services, automate workflows, or achieve networked programmability. A new paradigm is needed that leverages the existing biotech ecosystem to create PCLs that are useful, networked, & programmable.
Nexus Bio: An Interconnected Network of Programmable Cloud Laboratories to Empower U.S. Biomanufacturing
Nexus Bio is a cloud-based platform that converts existing service providers into a PCL and connects all PCLs together into a Programmable Cloud Network, streamlining the mesh of interactions between researchers and service providers. We are designing Nexus Bio to cultivate a thriving ecosystem: a vast community of service providers with thousands of on-boarded workflows who find the network useful because it enables frictionless interactions with researchers, increasing their revenue and reducing their operational costs; and a vast community of researchers carrying out thousands of experiments, who find the network useful because it enables them to formulate & test hypotheses, discover fundamental principles, and develop commercial products with a speed & scale that would be impossible for any single laboratory.
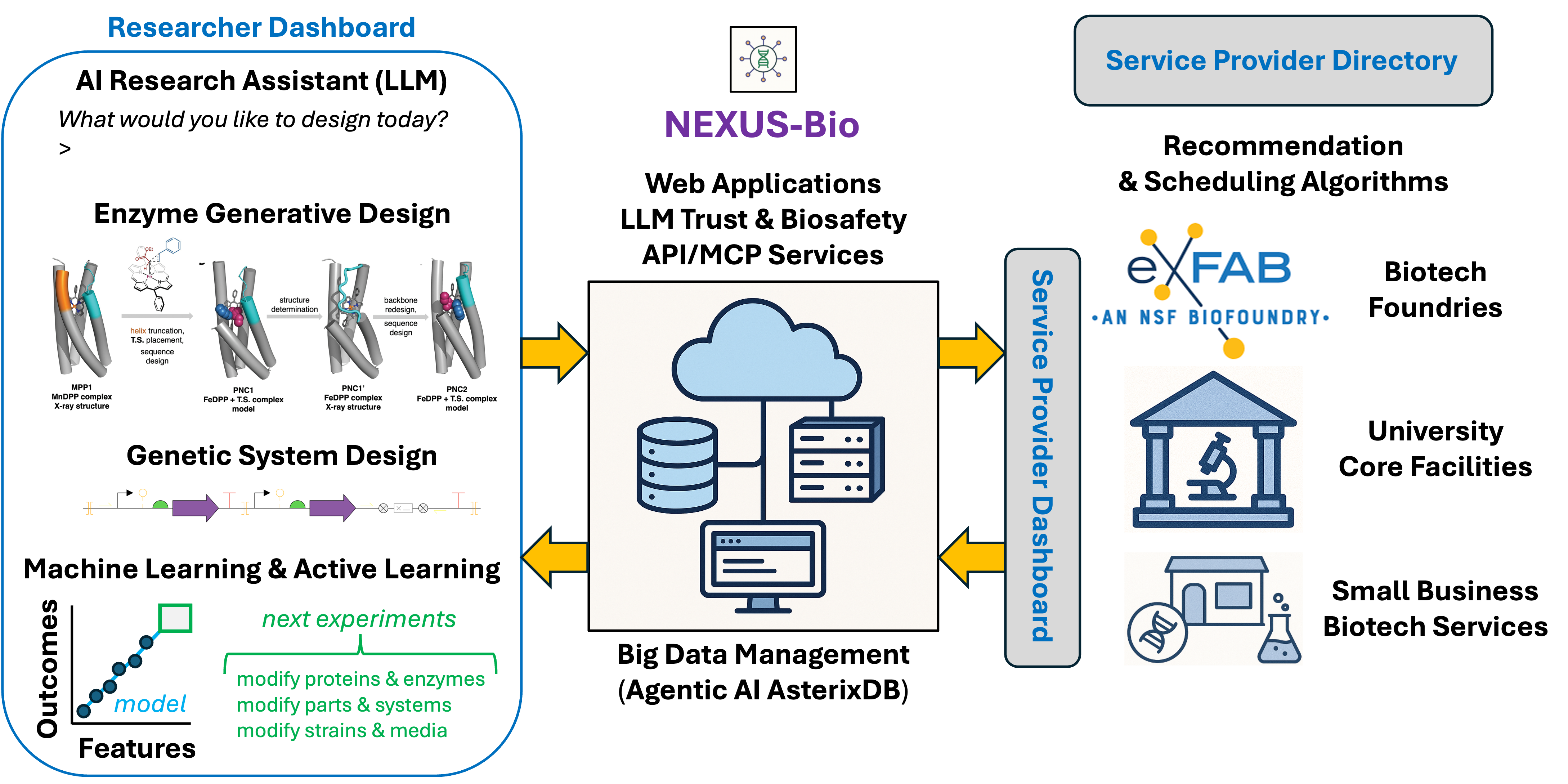
Nexus Bio will massively accelerate the speed and scale of biotechnology research
Nexus Bio for Researchers
Designs and executes a massive variety of complex, multi-step experiments according to your high-level specifications. Unlocks low-cost access to expensive, specialized equipment run by best-in-class biofoundries, core facilities, and commercial services. Provides end-to-end project management, including biosafety checks, workflow chaining, provider selection, scheduling, logistics, ML-ready data management, data visualization, and export. Harnesses AI research assistants, automated design algorithms, active learning, and scheduling optimization to massively expand your hypothesis testing and accelerate your design-build-test-learn cycle.
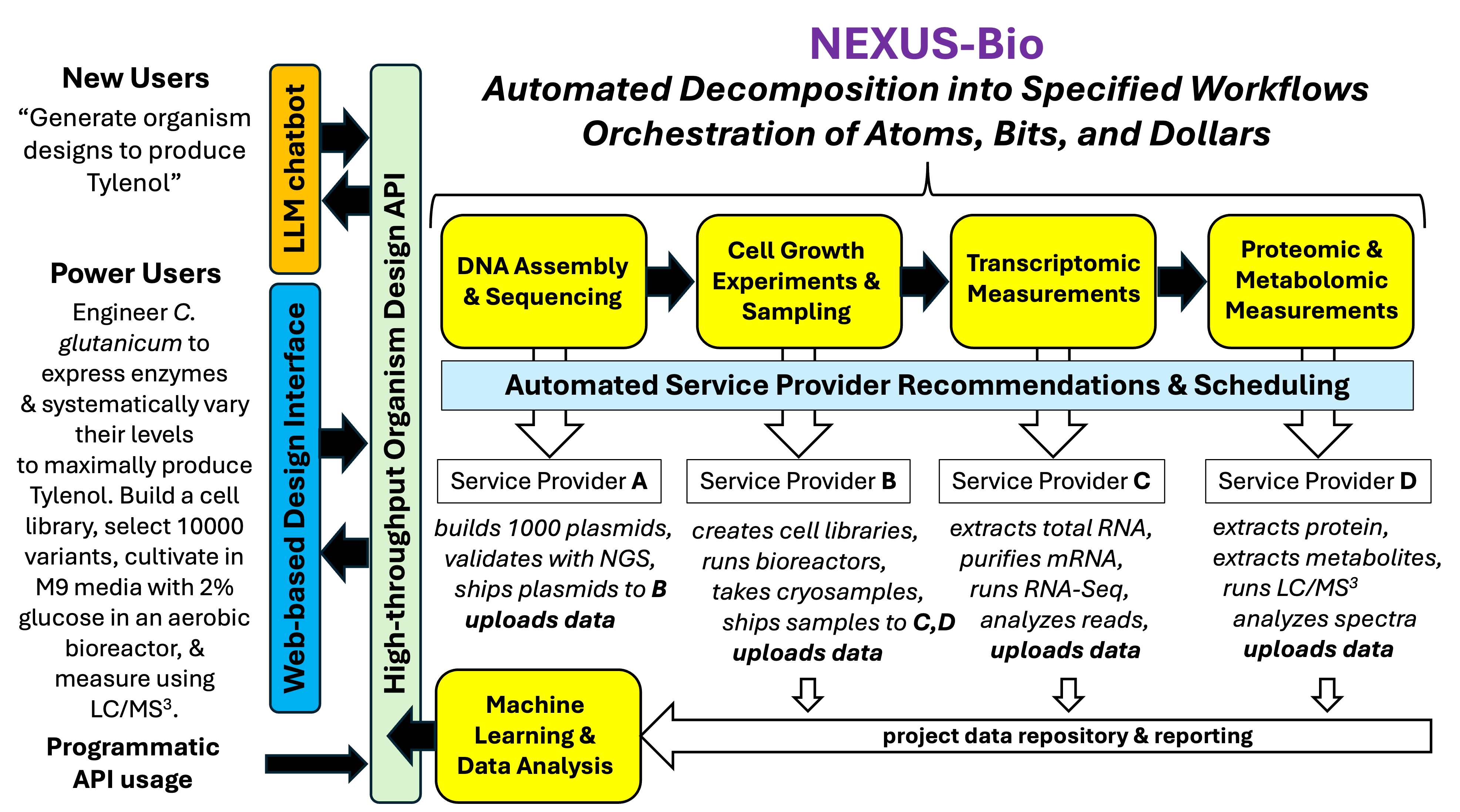
Nexus Bio for University Core Facilities
Expands research impact by attracting external academic and industrial users through a trusted national network. Increases utilization of your staff expertise and instruments by streamlining project intake, scheduling, data transfer, billing, and reporting. Provides external revenue while reducing your administrative overhead via easy researcher on-boarding, project tracking, and automated payments. Become AI-ready by converting your existing workflows into networked, programmable services without requiring major automation investments.

Nexus Bio for Commercial Service Providers
Advertises your services to a large and growing researcher base. Lowers customer acquisition costs by automatically scheduling researcher projects with your service workflows. Increases revenue through higher utilization, repeat customers, and participation in multi-provider pipelines. Simplifies operations via standardized project specifications, logistics coordination, automated payments, and data delivery. Improves your branding by showcasing your performance, turnaround time, and quality, rather than competing solely on price.

Nexus Bio will ...
- on-board researchers once for all networked service providers - reduces costly administrative overhead from O(N) to O(1)!
- provide a real-time directory of service provider workflows, pricing, and availability - advertises state-of-the-art workflows to all researchers with reviews
- design thousands of experiments for researchers - combining agentic AI with automated design algorithms for engineering pharmaceuticals, proteins, pathways, sensors, cells, & more
- break down complex experiments into well-defined chains of service provider workflows - automatically orders & ships custom materials (DNA, strains) to service providers
- schedule workflows with specific service providers, minimizing overall cost and turnaround time - maximizes all-network utilization & all-researcher satisfaction
- provide instructions to researchers and service providers with easy-to-use web-based dashboards - eliminates miscommunication and missing information
- track physical shipments and coordinate data transfer between researchers & service providers - creates a ML-ready data & metadata repository for every project
- orchestrate enterprise payment flows from researchers across to multiple service providers - one purchase order to pay many service providers
- accelerate data analysis by providing machine learning and data visualization tools - easy when all data & metadata is ready for ML analysis
- design the next round of experiments for researchers using active learning - suggests new designs & experimental conditions to maximize the desired objective
- prevent biosafety hazards with digital and physical layers of security - provides chemical and biological safety data to researchers & service providers, while preventing unauthorized use
- protect intellectual property and confidential information using the NIST 800-53 cybersecurity standard - service providers only receive the minimum data needed to carry out workflows
The Nexus Bio Team

















Meet the Companies & Core Facilities Powering the Nexus Bio Network v1.0
Join us!
Our Vision of an Internet of Programmable Cloud Laboratories
Nexus Bio in Action
Denise N Adams is a biotech researcher engineering an organism that will over-produce a glucagon-like peptide 1 (GLP1) inhibitor [exendin-(9–39)-NH₂], commonly used to treat diabetes. She logs into the Nexus Bio web-based platform using her institutional login & pre-arranged payment methods. Denise selects her favorite AI Research Assistant, and after a brief chat, returns with a textual description of the key ingredients: a modified pre-pro form of exendin with a yeast secretion tag, a secreted endoprotease (Kex2) that internally cleaves exendin, secreted exopeptidases (Kex1/Ste13) to trim the pro-peptide, and a secreted peptidylglycine α-amidating monooxygenase (PAM) to amidate the C-terminus & produce the active product. Nexus Bio retrieves these proteins’ amino acid sequences from an authoritative database (Uniprot) and stores all this specification data in a standardized project data structure.
Denise then selects her favorite Organism Designer from a Nexus Bio service provider, which identifies the optimal fungal host to express these enzymes (Saccharomyces cerevisiae uSEC16), selects combinations of genetic parts to express these proteins at different levels, and generates a list of 300 genetic system sequences. Denise uses the Nexus Bio New Project Interface to review the designed organisms and approve the project for building & testing.
Following the design workflow, Nexus Bio automatically queries its Service Provider Directory and compiles a list of biofoundries, core facilities, and commercial vendors that offer DNA synthesis & assembly, strain engineering, next-generation sequencing, proteomics, & other services together with their pricing, turnaround times, and track records of success. Nexus Bio leverages its Recommendation and Scheduling Algorithms to propose competitively priced, reliable service providers that all have adequate availability and throughput. Denise selects workflows from a biofoundry, a genomics core facility, and a bioinformatics core. They each receive New Project Notifications on their Service Provider Dashboards listing the relevant project data and the requested workflows. Each service provider reviews their project data, confirms their availability, and clicks the “Accept Project” button.
Nexus Bio orders the fungal strain from an authoritative strain repository (ATCC) and purchases the DNA building materials, both shipped to the biofoundry directly to lower biosafety risk. The biofoundry receives the shipment & workflow data on their dashboard. They construct 300 engineered Saccharomyces cerevisiae uSEC16 strains that produce the GLP1 inhibitor at varying production rates. They create cryostocks, send genomic DNA samples to the genomics core facility, and update the project data on their dashboard. The genomics core facility also receives the workflow data from their dashboard. They carry out the sequencing workflow on the engineered strains and upload the data onto their dashboard. The bioinformatics core then receives a notification that the sequencing data is ready; after their computational pipeline is completed, the bioinformatics core uploads the genome sequences back onto their dashboard. With every step, Denise receives an update on her project dashboard, showing progress, completion, and Interactive Data Visualization & Export for each data type.
Denise finds that 95% of the engineered strains match the expected designs (success!). She selects them for small-scale cultivation and characterization, customizing a proteomics workflow to identify the secreted peptide products, measure the amount of GLP1 inhibitor in supernatant, and quantify the different types of post-translational modifications. From its up-to-date directory, Nexus Bio finds a proteomics core facility with the needed instruments & availability, sending a New Project Notification to its dashboard. Nexus Bio sends another project notification to the biofoundry to grow the 285 strains in induced small-scale cultures and ship supernatant samples to the proteomics core facility. Nexus Bio also orders a positive control sample (chemically synthesized GLP1 inhibitor) and ships it directly to the proteomics core facility. The proteomics core facility receives the positive control & samples and runs the same LC/MS3 workflow on them, followed by analysis and data upload onto its dashboard.
Denise receives the raw & analyzed data via her project dashboard, showing that the top strain produced & secreted over 1 g/L of GLP1 inhibitor with 65% containing the correct modifications (a great start!). She leverages the Nexus Bio’s toolbox of Machine Learning algorithms to identify an important pattern in her data: insufficient PAM activity was responsible for 30% of the incorrect peptide product. This pattern is fed to the AI assistant, which reminds Denise that the PAM enzyme requires copper and ascorbate for optimal activity; the AI assistant also suggests to anchor the PAM enzyme in the membrane to amidate the peptide as it is being secreted.
Denise selects Nexus Bio’s Active Learning service, which redesigns the genetic system and modifies the media formulation. After reviewing the designs, Denise sends the new project to Nexus Bio for completion. Nexus Bio schedules the workflows as before, re-using the same service providers to maximize project knowledge & re-use of materials. The results come back in 3 weeks; GLP1 production is now 2 g/L with 95% active product. Delighted, Denise sends a notification to retrieve the engineered strains & materials, shipped to her work address. For her next project, Denise will optimize the media formulation and scale-up production & purification.
With Nexus Bio, Denise powers through cycles of organism engineering & product development, all without over-spending on expensive equipment. Nexus Bio enables Denise – the CEO of a 1-year start-up biotech company – to rapidly innovate, operate, and succeed. Every year, more service providers join Nexus Bio and further expand its workflows. New AI assistants and interconnected applications are built on top of Nexus Bio’s open data structures, APIs, and algorithms, enabling Nexus Bio to provide a portfolio of interconnected services that no single company – even a large pharmaceutical – could hope to replicate.
The Internet of Programmable Cloud Laboratories has arrived.
Our Progress
$20M Proposal Submitted to National Science Foundation
The Nexus Bio concept was developed in response to the National Science Foundation's ambitious and innovative PCL-Testbed program. Our inter-disciplinary team proposed to develop novel algorithms and software systems to implement our innovative Nexus Bio network. We will test its capabilities on specific projects using an initial network of 10 service providers providing over 200 interconnected biotech workflows.
Work with Us
Tell us how you’d like to engage, and we’ll follow up.